Crosstalk of cell shape and state
Stochastic Cell shape dynamics during epithelial-to-mesenchymal transition
Further information will be added soon
Morphometric analysis of Amphioxus notochord cell shapes
Further information will be added soon
Morphodynamics of insect gastrulation
Further information will be added soon
Aggregation Processes
Irreversible protein aggregation in the presence of phase-separated droplets
Further information will be added soon
Assembly of cellular aggregates
Type IV pili can mediate attractive interactions between bacterial cells and the motion of those cells on top of a substrate. Together with Dr. Vasily Zaburdaev and Dr. Christoph Weber, I developed a mathematical model to study how aggregation (resulting from the coalescence of microcolonies moving on the substrate), fragmentation (the loss of single cells from colonies) and proliferation affects the colony size density.

We compare our theoretical results to experimental data, contributed by Prof. Nicolas Biais, and observe excellent agreement.
Cell Motility of prokaryotes and eukaryotes
Motility of single bacteria and bacterial colonies on a substrate
Next to attractive cell-cell-interactions, type IV pili also mediate the motion of bacteria on a substrate by a mechanism similar to a grappling hook. Such substrate-associated motion is observed for a wide range of bacterial cells, for example, Pseudomonas aeruginosa, Neisseria meningitidis or Myxococcus xanthus. For Neisseria gonorrhoeae, it was previously reported that the bacteria exhibit a persistent random motion on glass surfaces, with a characteristic length scale that exceeds the length of pili. In the group of Dr. Vasily Zaburdaev, I develop theoretical and numerical models to unravel how interactions of pili can mediate such peculiar behavior. Opposing to previous reports, the collective behavior of multiple pili alone is sufficient to explain the persistence of the motion, without the need to consider any directional memory. We also find that sliding friction has a huge impact on the motility of microcolonies.
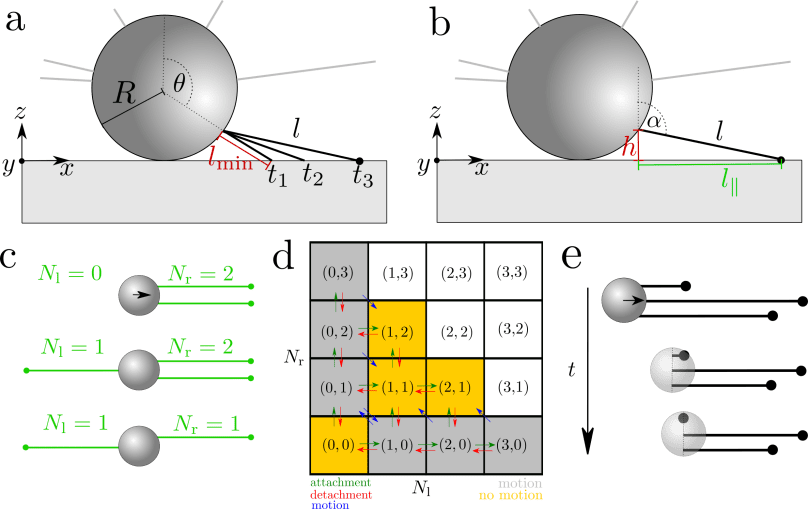
Publications: Pönisch et al., NJP 2017, Pönisch et al., PRE 2019
Motility of stem cells in confinement
Further information will be added soon
Dynamics of bacterial aggregates
Internal dynamics of bacterial microcolonies and colony coalescence
In order to considerably increase their survivability, bacteria from aggregates so-called microcolonies. These colonies are the precursor of biofilms, surface-associated communities of bacteria that are embedded in an exopolysaccharide matrix. While commonly one imagines bacteria as individual entities, the majority of bacteria is associated with such microcolonies and biofilms. Bacteria form microcolonies with the help of type IV pili, long and thin appendages that can reach lengths exceeding the cell size. The pili can mediate attractive cell-cell-interactions by cycles of protrusion, binding to other pili and retraction.

I study how exactly pili can mediate the dynamics of bacterial microcolonies. Specifically, we investigate experimentally (in collaboration with Prof. Nicolas Biais) and with the help of theoretical modeling (together with Dr. Vasily Zaburdaev) the coalescence of two colonies of Neisseria gonorrhoeae microcolonies, which exhibits behavior that differs from the coalescence of fluid droplets. We could explain this observation by a spatially-dependent gradient of motility of individual cells within microcolonies.

Additionally, I apply numerical tools to study how mixtures of wildtype cells and different cells with mutations of their pilus apparatus demix. For example, a mixture of cells with pili that are not capable of retraction and thus are not able to mediate an attractive cell-cell-force and of wildtype cells exhibits demixing. The mutant cells (red in the image directly above) are located at the surface of colonies, wildtype cells (green) within the colony bulk.
Publications: Pönisch et al., NJP 2017, Pönisch et al., Scientific Reports 2018 and Pönisch et al., EPJ B 2018