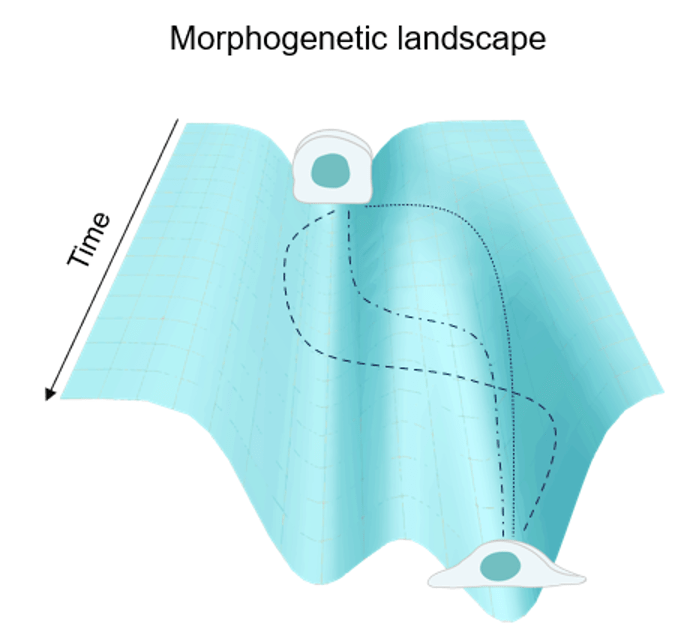
My main postdoc work is finally out as a preprint! In this study, we investigate whether cell shape changes during cell state transitions can be described as a particle moving in a morphogenetic landscape, similar to gene expression trajectories in a Waddington landscape. We were super excited when we found that this is not only possible when describing epithelial-to-mesenchymal transition (EMT), but that cell shape changes during EMT are associated with a transient noise peak! A combination of mathematical modelling and experiments showed that this noise peak is robustly accelerating shape changes during EMT! Interested, have a look!
This has been a great collaboration with my awesome co-first author Iskra Yanakieva and together with Ewa Paluch and Guillaume Salbreux.